Plant–pathogen interactions have been studied mainly in agricultural crops with important applications in the selection of disease-resistant cultivars and in improving the understanding of the molecular control of gene regulation. But relatively little work has been done on tree pathosystems. Exploration of the literature on several aspects of investigations on forest pathology also testifies to the fact that basic research on the study of diseases of forest trees lags far behind parallel work on agricultural crop pathology.

(Illustration: Jan Stenlid)
The knowledge about the biochemical and molecular aspects of host-parasite interaction in tree pathosystem is considered critical for a better understanding of variation in natural resistance and also as a basis for increasing resistance through breeding and or genetic engineering.

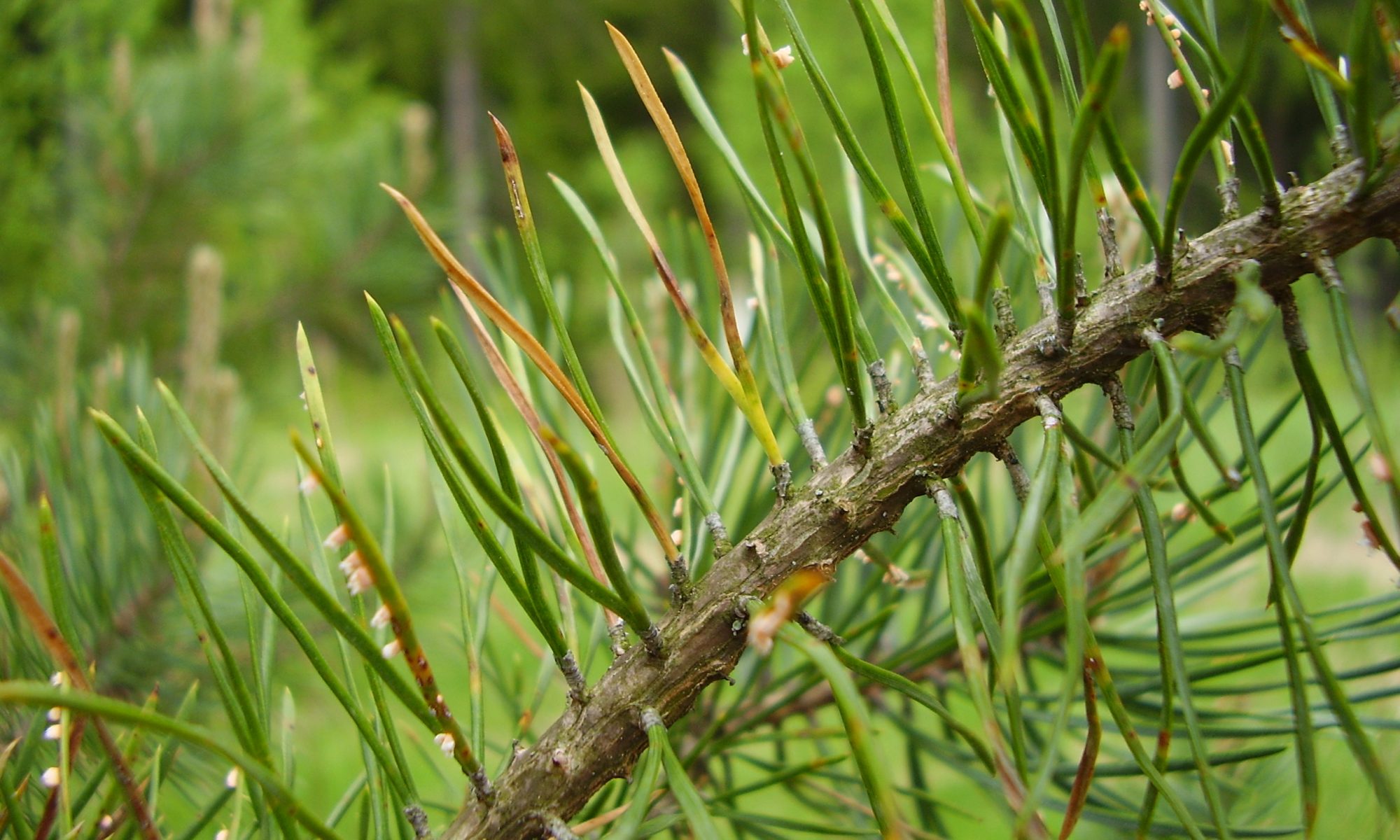
In tree pathosystems, H. annosum root rot (lFig. 1a-b) has been the focus of several studies (Asiegbu et al. 2005). Over the years, our primary mission has been to integrate both basic, applied and descriptive research in forest pathology with the hope that the information obtained will provide new insights into cellular processes as well as lead to the development of novel forest protection and fungal based products for tackling forest tree diseases.
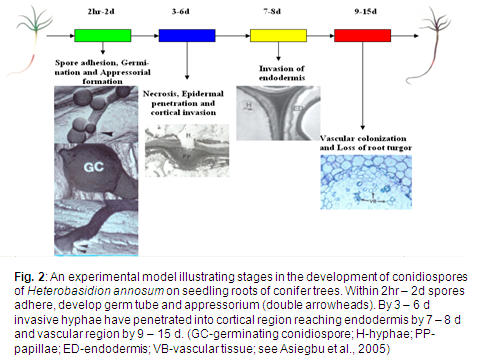
Our major research focus has been the determination of the genes as well as cellular (Fig. 2), physiological, molecular and biochemical pathways required by forest pathogens to infect and cause disease to conifer trees.
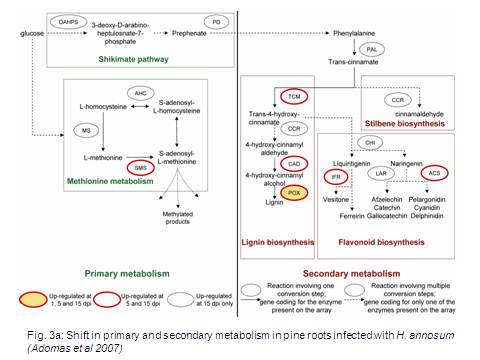
Equally, elucidating host factors (Fig. 3a) which confer resistance against necrotrophic parasites like H. annosum is also a primary research goal. An additional long term goal is to understand how forest trees are able to recognise some fungi as pathogen whereas they are able to form intimate beneficial association with others.
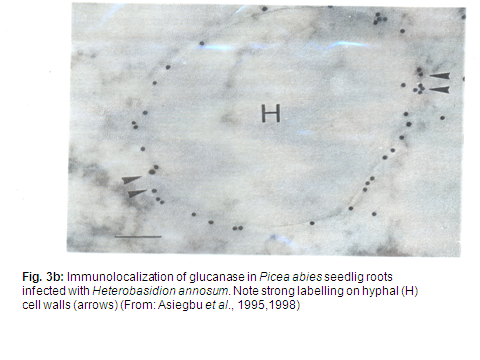
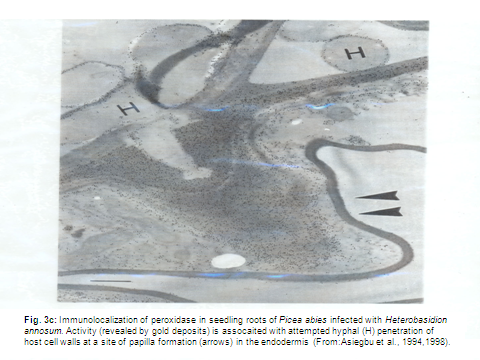
In exploring the differential basis for resistance of conifer tissues challenged with either parasitic or non-pathogenic microbes, we study both immature juvenile conifer seedlings and mature differentiated living wood-xylem tissues, utilizing widely diverse techniques and approaches including; transcript profiling, comparative genomics, structural and functional biology, HPLC, radioisotpes, lectins, fluoresence microscopy, antibodies, ELISA, scanning and transmission electron microscopy, immunocytochemistry (Fig. 3b-c) and molecular methods.