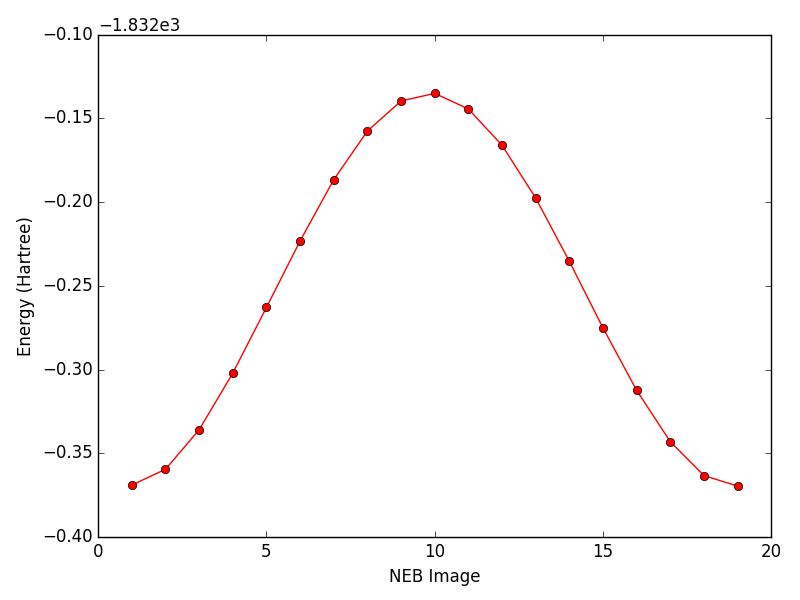
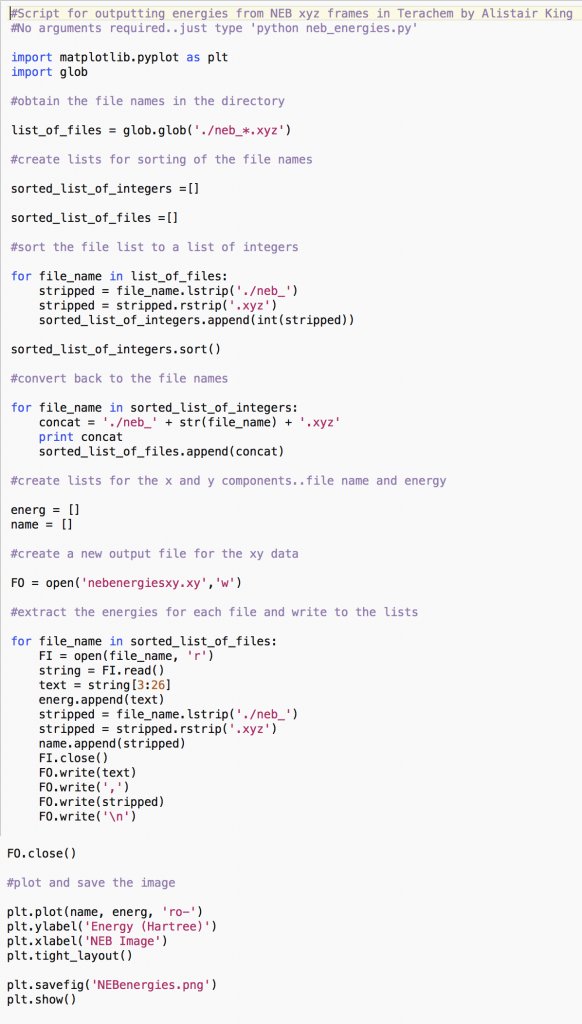
This python (python.org) script (neb_energies.py) can be used for parsing the neb_*.xyz ouput geometries (where * is the geometry frame number) from a nudged elastic band (NEB) ‘transition-state’ search in Terachem (petachem.com). It extracts the energies (in Hartrees) for each frame from the xyz files and plots them using matplotlib (matplotlib.org) for quick viewing.
Category Archives: Scripting
Visualisation of Energies from Geometry Optimisations using TeraChem
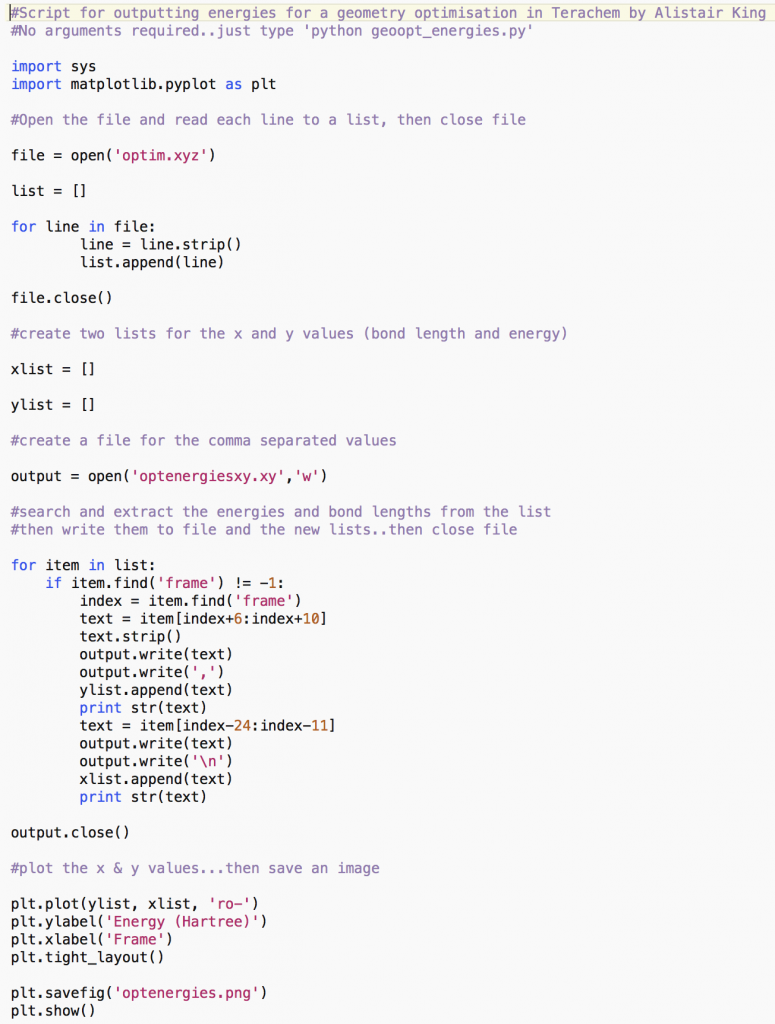
This python (python.org) script (geoopt_energies.py) can be used for parsing the geometry.xyz ouput geometries (typically optim.xyz) from a geometry optimisation in Terachem (petachem.com). It extracts the energies (in Hartrees) for each frame from the xyz file and plots them using matplotlib (matplotlib.org), for quick viewing.
Visualisation of Energies from a Potential Energy Surface (PES) Scan using Terachem
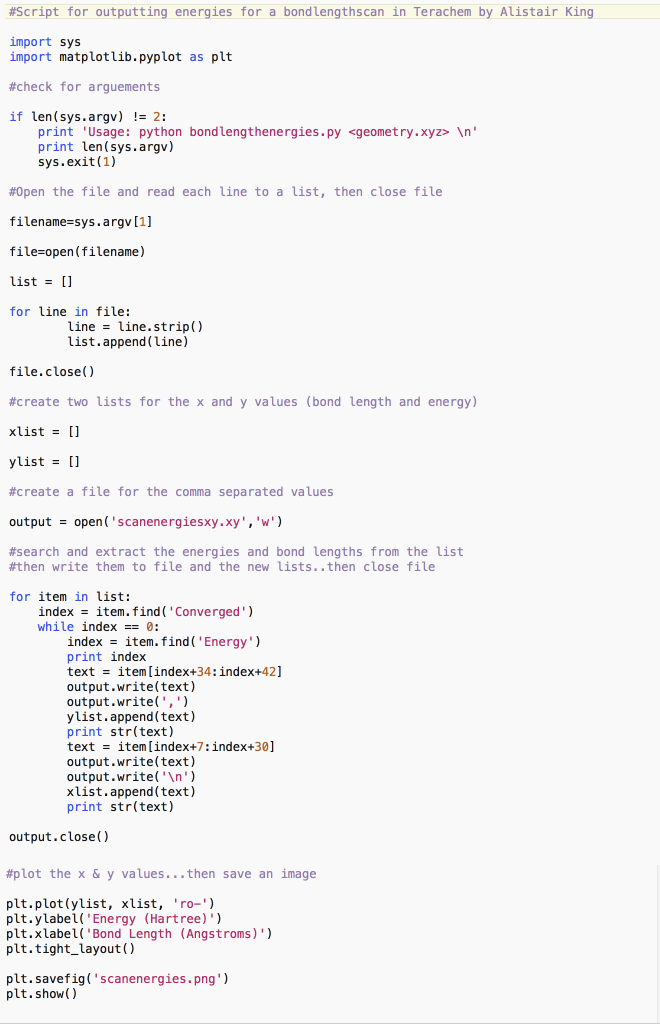
This python (python.org) script (bondlength_energies.py) can be used for parsing the geometry.xyz ouput geometries (typically optim.xyz or scan_optim.xyz) from a potential energy surface search in Terachem (petachem.com). This version, by title only, is used to extract the enegies for bond-length searches but is equally suited for variation in other geometry features, e.g. bond angles. It extracts the energies (in Hartrees) for each frame from the xyz file and plots them using matplotlib (matplotlib.org), for quick viewing.
Proton Affinitiy Calculation from GAMESS Outputs
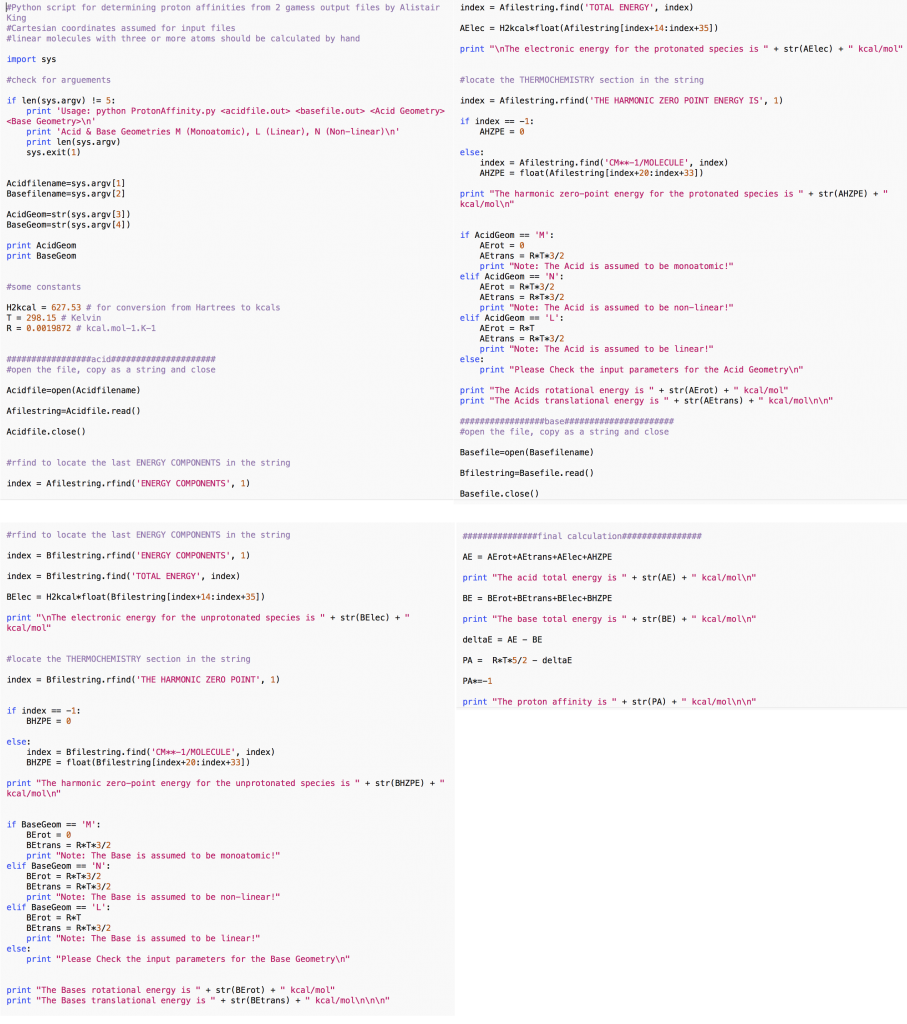
This script is used for rapid calculation of proton affinity values (enthalpy of separation of a proton to affinity, typically in the gas-phase) from GAMESS outputs.
Essentially the energies of the protonated vs the unprotonated species are calculated and subtracted to give the energy value. The calculation was based on a summer school tutorial organised by the Theoretical and Computational Biophysics Group from the University of Illinois at Urbana-Champaign but also after discussion with Dr. Gordon Driver.
The script was published in a recent article on the prediction of cellulose dissolution capabilties of acid-based conjugate ionic liquids.
DS Determination from 31P NMR
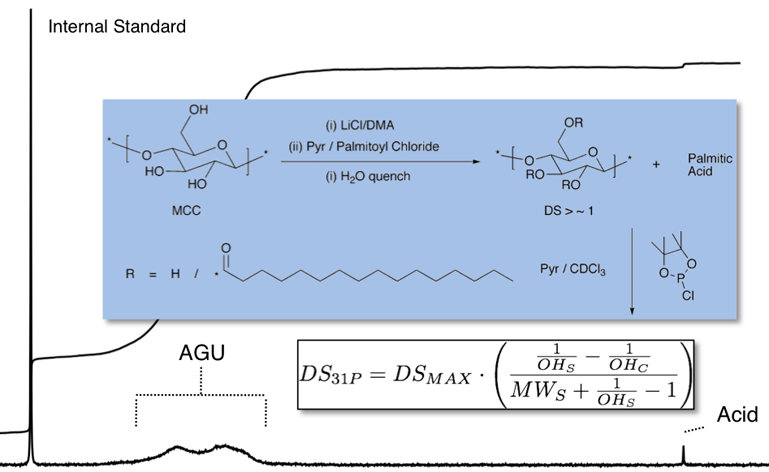
Another script that I wrote to determine degrees of subsitution (DS) on modified cellulose was based on 31P NMR analysis.
Chemical modified celluloses are derivatised as phosphite esters (Fig). Those phosphite esters are quantified against an internal standard to allow for the determination of the ‘available hydroxyls’ per mass unit of sample. Based on the molecular weight of the additional functionality and the available hydroxyls value the DS can be calculated.
This script and the method development were published in the journal Analytical Methods.
DS Determination from Elemental Analysis
The first python script that I wrote was to determine degrees of subsitution (DS) on modified cellulose from elemental analysis results. Details of the different atoms added to cellulose and the elemental analysis outputs for C, H, O, N, S.
DS values are calculated for the different elements in the sample. Naturally the calculation based on hydrogen gives significant error. Carbon is a little better and if the sample contains nitrogen the results will be most accurate.
The script also contains the whole periodic table in the python ‘dictionary’ format.
The script works by calculation of the different elemental compositions for iterative increases in DS value. The values are then compared to the actual experimental elemental analysis values to determine the DS values for the diferent elements.